Bioinformatics
Pipelines
Automated, validated analysis from raw sequencing data to structured outputs. Run any nf-core pipeline or import your own from GitHub, without writing infrastructure code.
The Problem
Genomic analysis shouldn't break every time you change a dataset, a collaborator, or a question.
Most teams still stitch together fragmented tools and ad-hoc scripts. The result: workflows that can't be repeated, compared, or scaled.
Fragmented tooling
Every project reinvents the wheel, custom scripts, local installs, undocumented pipelines. Nothing connects, nothing carries over.
Slow time-to-insight
Weeks spent debugging environments and reformatting outputs instead of interpreting biology. The bottleneck is infrastructure, not science.
Can't scale across teams
What works on one machine breaks on another. Results vary between analysts. Collaboration means starting from scratch.
How It Works
From raw data to results
in five steps
A streamlined path from sequencing files to clinical-grade outputs, automated, reproducible, and auditable.
sample_R1.fastq.gz
2.4 GB
sample_R2.fastq.gz
2.4 GB
samplesheet.csv
12 KB
Step 01
Upload sequencing data in standard formats, FASTQ files, sample sheets in CSV or TSV, and more.
-
Step 01: Upload Your Data
Upload sequencing data in standard formats, FASTQ files, sample sheets in CSV or TSV, and more.
-
Step 02: Choose Your Analysis
Select from validated nf-core pipelines or import your own Nextflow workflow directly from GitHub.
-
Step 03: Pipeline Processes
The pipeline handles quality checks, data alignment, variant or expression analysis, and preparation of structured outputs.
-
Step 04: Explore in Notebooks
Walk through results step by step in interactive notebooks, adjust thresholds, run custom analyses, and visualize findings.
-
Step 05: Outputs & Reports
Receive structured outputs ready for interpretation, tables, figures, VCFs, BAMs, and clinical reports.
Automated Pipelines
Validated workflows.
Consistent results.
Every pipeline is organized, versioned, and production-ready, so your team can focus on biology, not infrastructure.
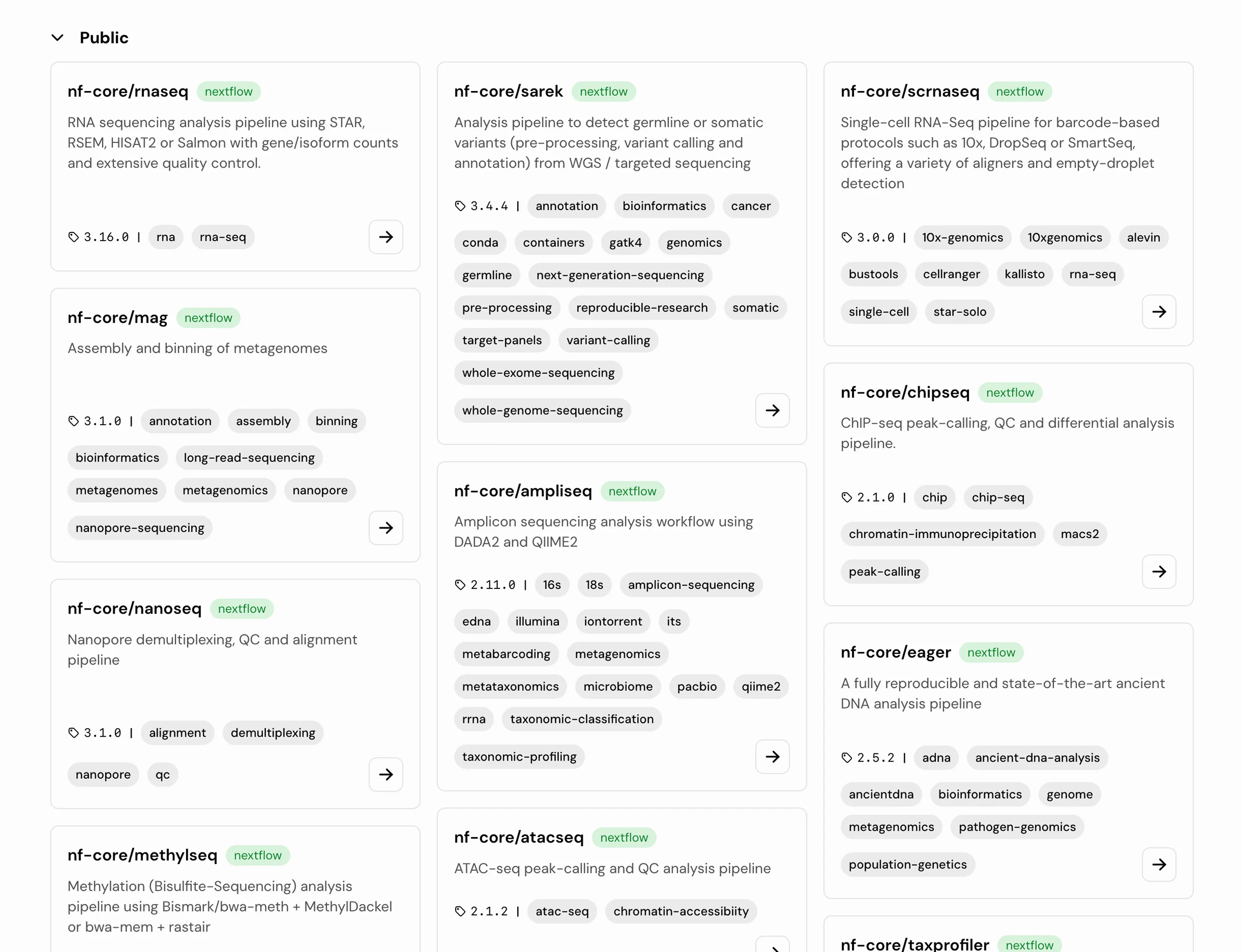
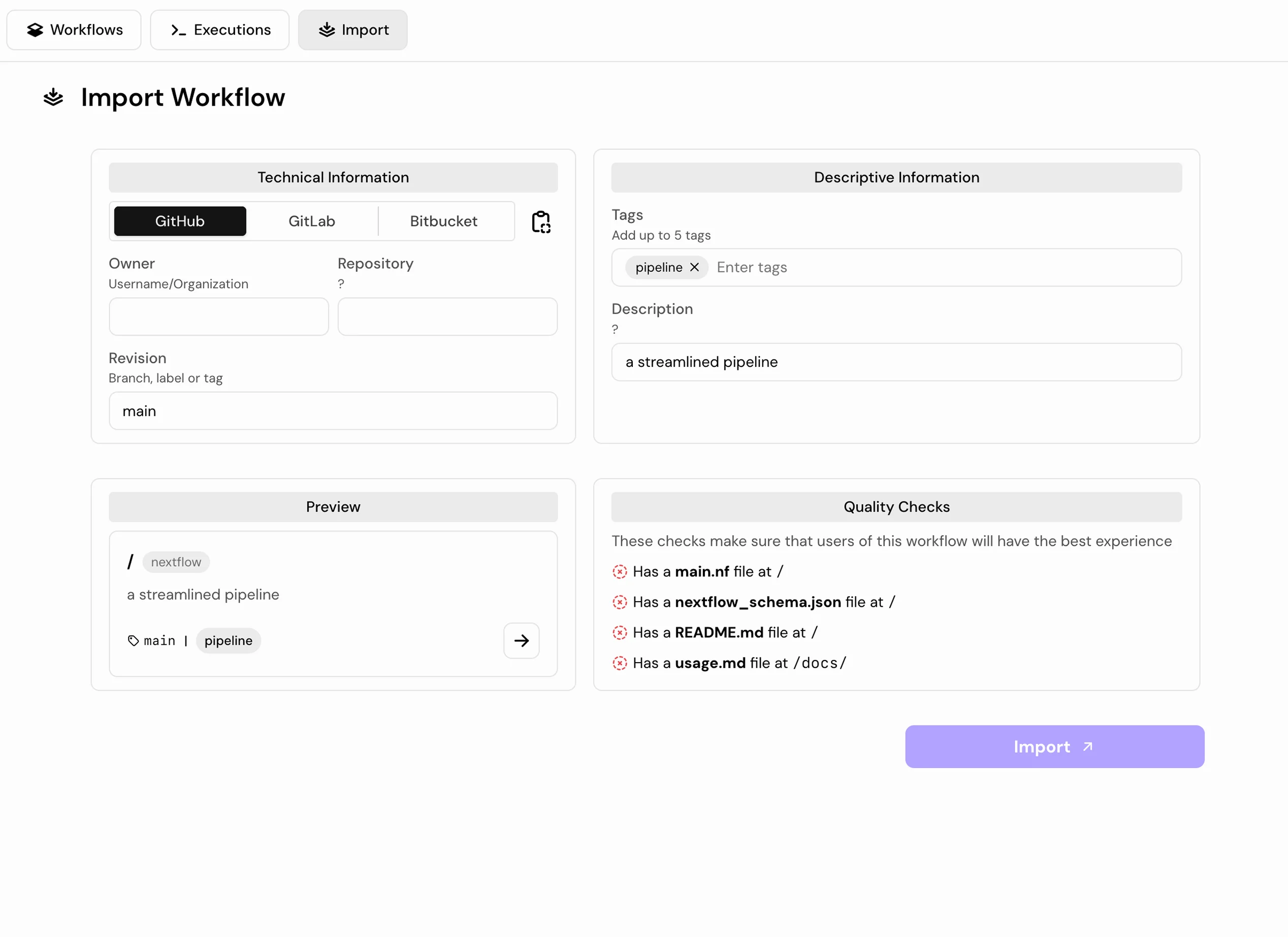
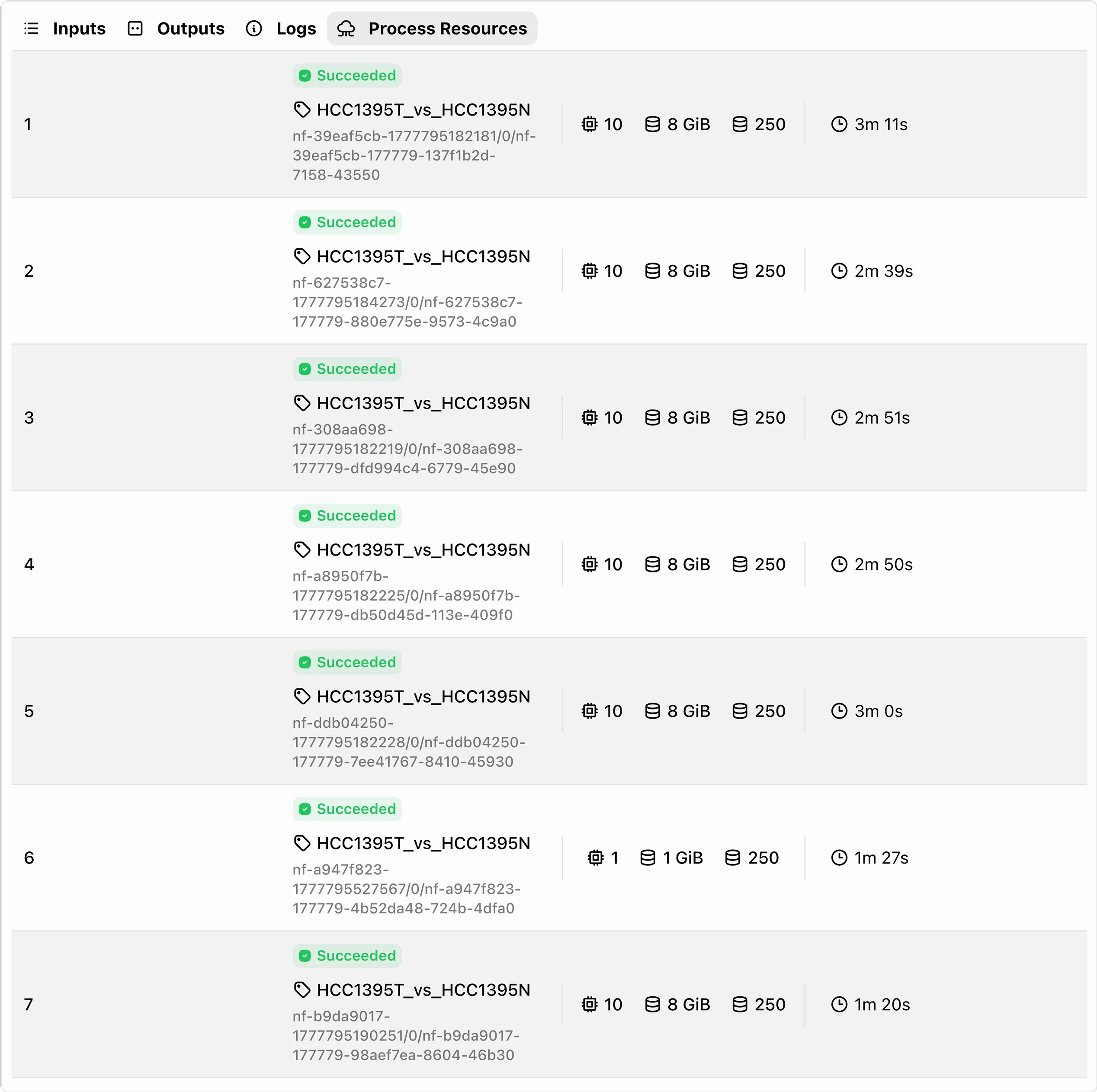
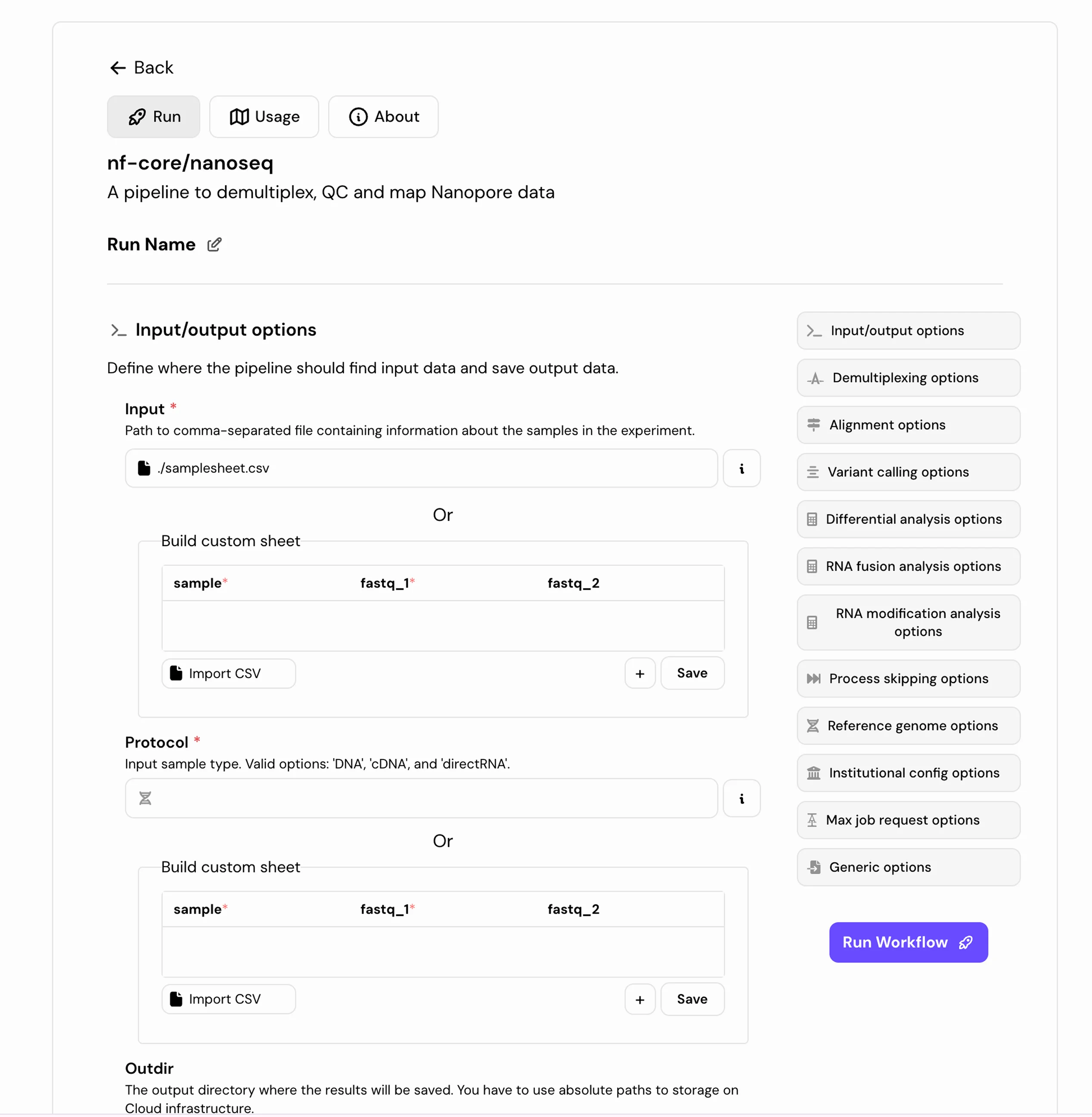
AI-Assisted Pipelines
Powered by BioMind.
Guided by your data.
The Pipeline Agent works alongside every stage of your analysis, from selecting the right workflow to explaining why a run failed.
BioMind
AI Your AI pipeline guide,
for every stage of the run.
Pan.bio's Pipeline Agent has deep knowledge of every nf-core pipeline it supports. It guides parameter selection for your sequencing protocol, validates your sample sheet before submission, and explains outputs in plain language — so your team spends less time reading docs and more time interpreting results.
Parameter Guidance
Select the right nf-core pipeline and configure run parameters for your sequencing protocol, genome, and variant calling mode before you start.
Documentation on Demand
Ask questions about any pipeline step and get answers directly from nf-core documentation, with the relevant section cited.
Output Interpretation
Explain MultiQC metrics, identify quality flags, and suggest the right downstream steps based on what your pipeline actually produced.
Failure Diagnosis
Read pipeline failure logs, explain what went wrong in plain language, and suggest the right recovery path.
Built on nf-core Knowledge
Unlike a generic code assistant, BioMind's Pipeline Agent is trained on nf-core pipeline documentation, common parameter pitfalls, and protocol-specific configurations. Ask it about Sarek germline mode vs. tumor-normal, or why your RNA-seq run is flagging a low alignment rate, and it responds with pipeline-specific guidance — not a generic answer.
Collaboration
Built for Teams,
Not Just Individuals.
Powerful tools mean nothing if your team can't standardize on them. Pan.bio is designed so that every pipeline, every notebook, and every analysis is structured for collaboration, reproducible by default, auditable by design, and accessible to everyone on the team.
Shared Notebooks
Multiple users collaborate in the same notebook, sharing data and analysis while each maintaining their own interaction history with BioMind.
Reproducible Standards
Pipelines are always available with specified versions. Successful experiments become shared multi-step analyses your entire team can standardize on.
Audit Trail & Logging
Every analysis is logged, versioned, and auditable. Know who ran what, when, and with which parameters, across every project.
Access Control
Role-based permissions and multi-tenant isolation ensure teams work securely. Each client or project stays separated without rebuilding the stack.
For Researchers
Pipelines reduce complexity and save time. Instead of worrying about software setups or technical details, receive structured outputs that are ready for interpretation or reporting. Analyses become easier to repeat, compare, and scale across projects.
For Organizations
Workflows become consistent, easier to maintain, and faster to deploy. Teams collaborate more effectively because everyone works from the same structured process rather than individual scripts or ad-hoc methods.
Differentiators
What Makes Pan.bio
Workflows Different?
What sets Pan.bio apart is the balance between structure and flexibility. Many platforms offer automation but limit exploration. Others allow full flexibility but become difficult to maintain at scale. Pan.bio connects the two, helping teams move faster without sacrificing clarity or trust in the results.
Automation Where It Matters
Reliable, repeatable processing through validated, version-controlled pipelines.
Notebooks for Deeper Exploration
Pipeline outputs flow directly into Notebooks for interactive analysis, no re-uploading, no context-switching.
Reusable by Design
Turn successful experiments into shared multi-step analyses your entire team can standardize on.
AI-Powered Assistance
BioMind guides parameter configuration and answers questions about any pipeline, cited from its own documentation.
Designed for Real-World Use
Built to handle both research and production environments with enterprise-grade compliance.
Enforced at the infrastructure level, your data stays in your jurisdiction, always.
See It in Action
Pan.bio Bioinformatics Pipelines turns fragmented genomic analysis into a structured, reproducible workflow, from raw sequencing data to structured outputs, every time.
No credit card required · Start in minutes